This project aims to review and improve current state of the art Neural-ODE implementations to dynamically predict patient Pharmacokinetics.
Current implementations have shown that Neural-ODE is the most accurate PK model in predicting untested treatment regimens and performs better than commonly used algorithms for time-series data such as LSTM and LGBM.
We will be reviewing and validating the results of the work done by past publications that implement Neural-ODE for PK/PD Modelling including:
- Neural-ODE for pharmacokinetics modeling and its advantage to alternative machine learning models in predicting new dosing regimens
- Deep learning prediction of patient response time course from early data via neural-pharmacokinetic/pharmacodynamic modelling
We will then conduct pressure-tests and bench-mark performance of different deep-learning methods to improve PK modelling architectures developed by publications.
- Phase 1 (Week 1-2): Read and understand the paper. Run the code and validate results in the paper.
- Phase 2 (Week 3-6): Translate codes from different sources into structured modules.
- Phase 3 (Week 7-8): Set up the PK/PD simulation through ODE equations to generate data.
- Phase 4 (Week 9-12): Conduct pressure test and bench-mark performance of different deep learning methods in plots.
| Column | Description |
|---|---|
NeuralODE_Paper_Supplementary_Code |
Supplementary Code from Paper 1 |
Supplement Code Mathematica |
Supplementary Code from Paper 2 |
Deep learning prediction of patient response time course from early data via neural-pharmacokinetic_pharmacodynamic modelling.pdf |
PDF of Paper 2 |
Neural ODE Reproduction.ipynb |
Reproduction of Code from Paper 1 in Jupyter Notebook |
Neural-ODE for PK.pdf |
PDF of Paper 1 |
Neuro-ODEs.pdf |
PDF of Pioneering Neural-ODEs Publication |
We want to standardize a protocol for branching during the course of this project so that we can avoid conflicts when merging and enable easier integration of changes into the main branch. This portion of the README.md will alo enhance productivity by ensuring proper coordination among our team and enable parallel development.
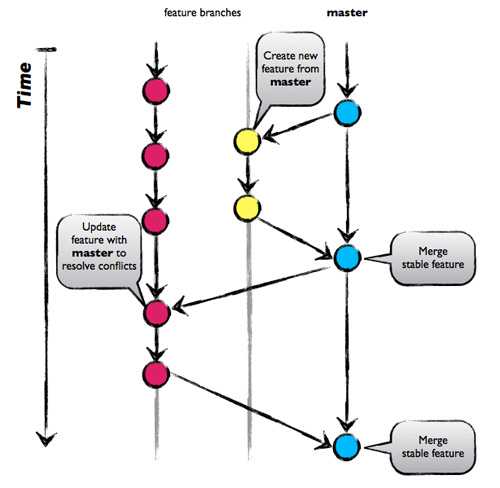
A visualization of the proposed strategy of branching is below:
This brancing strategy is known as Github Flow and it focuses on Agile principles and so it is a fast and streamlined branching strategy with short production cycles and frequent releases. Since there is no development branch as we are testing and automating changes to one branch which allows for quick and continuous deployment. This strategy is particularly suited for small teams such as ours since we need to maintain a single production version throughout.